- Témaindító
- #1
- Csatlakozás
- 2024.02.16.
- Üzenetek
- 753
- Reakció pontszám
- 39
- Kor
- 32
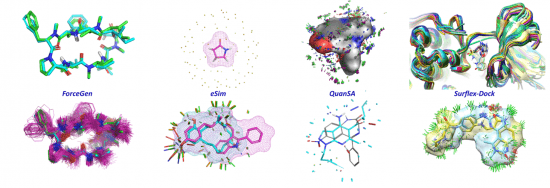
BioPharmics Surflex Platform 5.191 MultiOS
File Size: 2.8 GB
Tools Module
Fast and Accurate Small Molecule Processing
The Tools module addresses the most common aspects of small-molecule preparation
2D to 3D conversion (from SMILES or SDF)
Chirality detection and enumeration
Protonation
Conformer generation
Features and benefits
Template-free and non-stochastic
Relies on MMFF94sf forcefield for structure derivation
Fast and accurate on typical drug-like ligands, with better coverage of diverse conformations
Fastest and most accurate method for macrocyclic ligands
Capable of incorporating NMR restraints, which is particularly useful for large peptidic macrocycles
Similarity Module
State-of-the-Art 3D Molecular Similarity
The Similarity module implements ligand similarity operations using the eSim method
Virtual screening
Pose prediction
Multiple ligand alignment
The core eSim methodology is also integrated into the Docking and QuanSA modules.
Features and benefits
Virtual screening enrichment is both practically and statistically significantly better than alternative methods
Virtual screening speeds of over 20 million compounds per day on a single computing core
Databases of billions of molecules can be screened in hours using cloud-based computing resources
Pose prediction accuracy is substantially better than alternative approaches
Docking and xGen Modules
Top-Tier Solution for Virtual Screening and pose Prediction + Real-Space X-ray Density Modeling of Ligands
The Docking module addresses all aspects of ensemble docking
Large-scale PDB retrieval and processing
Surface-based binding site alignment using the PSIM method
Fully automatic pocket variant selection to cover the relevant protein conformational variation
Virtual screening
Pose prediction
Feature and benefits
Automated alignment and selection of appropriate binding site variants
Robust and fully automatic modes for virtual screening and pose prediction\Very extensive validation
Highly accurate non-cognate ligand docking
Directly applicable to synthetic macrocycles, with accuracy equivalent to non-macrocycles
The xGen module implements a novel method for real-space refinement and de novo fitting of ligand ensembles into X-ray density maps
Models ligand density using conformational ensembles
Avoids atom-specific B-factors as X-ray model parameters
Produces chemically sensible conformers with low strain energy; applicable to complex macrocycles
Yields superior fit to X-ray density than standard fitting approaches
Accessible to non-crystallographers and as part of crystallographic workflows
Affinity Module
Unique Machine-Learning Approach for Prediction Binding Affinity and Pose
The Affinity Module implements the QuanSA (Quantitative Surface-field Analysis) method, which builds physically meaningful models that approximate the causal basis of protein ligand interactions. The module implements integrated procedures for quantitative prediction of both binding affinity and ligand pose, with or without protein structural information
Multiple ligand alignment for molecular series that include multiple scaffolds
Incorporation of known binding site information
Machine-learning approach to physical binding site model induction using a multiple-instance approach
Prediction of both binding affinity and binding mode of new ligands
Iterative refinement of models with new data
Features and benefits
Fully automatic model building, including all aspects of ligand conformation and alignment
The binding site model (a "pocket-field") is analogous to a protein binding site, including aspects of flexibility
The pocket-field identifies which pose a new molecule must adopt, and ligand strain is directly modeled
Measurements of prediction confidence and molecular novelty guide user interpretation
Very detailed aspects of molecular surface shape, directional hydrogen bonding preferences, and Coulombic electrostatics are learned
Requires as few as 20 molecules for model induction and is capable of modeling series of hundreds of molecules
Please Buy Premium Account from my links to get high download speed and support me
https://rapidgator.net/folder/7686806/BioPharmicsSurflexPlatformv5191.html
https://nitroflare.com/view/A1C49DA...x.Platform.v5.191.MultiOS-BTCR.0624.part1.rar
https://nitroflare.com/view/BC00094...x.Platform.v5.191.MultiOS-BTCR.0624.part2.rar
https://rapidgator.net/file/095291a...tform.v5.191.MultiOS-BTCR.0624.part1.rar.html
https://rapidgator.net/file/1508a7f...tform.v5.191.MultiOS-BTCR.0624.part2.rar.html
[/code]
